Significance
Fourier ptychographic microscopy (FPM) has gradually taken shape as a practical way of sidestepping the familiar resolution–field-of-view compromise that limits standard wide-field microscopes. Instead of relying solely on hardware to push resolution, it constructs a synthetic high-NA aperture by illuminating the specimen from multiple angles. When this works well, the method produces surprisingly detailed reconstructions across expanses of tissue that would normally require tedious mechanical scanning. This capability has positioned FPM as an appealing tool for hematoxylin- and eosin-stained slides, where pathologists need both a broad overview and the ability to examine subtle morphological cues within the same dataset. However, the technique has never been entirely straightforward to implement and real systems don’t behave like the idealized models used in the forward formulation, and small deviations tend to accumulate in ways that undermine reconstruction quality. Moreover, variations in LED brightness, minor shifts in illumination direction, or the vignetting introduced by large illumination arrays all interfere with the model’s assumptions. Spatially varying aberrations further complicate matters, leaving traditional FPM pipelines heavily dependent on careful calibration and on raw measurements that are cleaner than what typical laboratory conditions deliver.
Most classical FPM algorithms frame reconstruction as a phase-retrieval task, alternating between spatial and Fourier constraints to recover both amplitude and phase. Although conceptually clear, these routines are slow and, more importantly, prone to stagnation when the data stray from the ideal forward model. Even more recent developments—such as feature-domain formulations that lean on image gradients—help with robustness but remain computationally demanding. Closed-form approaches offer a different perspective, but they come with strict requirements on illumination geometry and tend to falter when confronted with real-world artifacts like uneven illumination or residual vignetting. Taken together, these limitations point toward a lingering gap: the field still needs reconstruction strategies that can tolerate the imperfections of practical imaging hardware while preserving the resolution and stability required for large-area tissue microscopy. To this account, new research paper published in Photonics Research and conducted by Dr. Shuhe Zhang and Professor Liangcai Cao from the Tsinghua University alongside Miss Jiayun Li from the University of Oxford, researchers developed a latent-wavefront physical model for Fourier ptychographic microscopy, in which the unobserved camera-plane wavefront is explicitly treated as a latent variable. Using a variational EM algorithm, they alternated between estimating this latent wavefront and solving a closed-form deconvolution for the sample’s Fourier spectrum and pupil function.
The research team built and evaluated their VEM-FPM framework using a custom FPM platform equipped with a programmable LED array and low-magnification objective lenses designed to capture expansive tissue regions. Their data acquisition began with H&E-stained biological slides, illuminated sequentially by LEDs arranged in concentric rings above the sample. Each illumination angle produced a low-resolution intensity image, and together these measurements formed the backbone of the reconstruction problem.
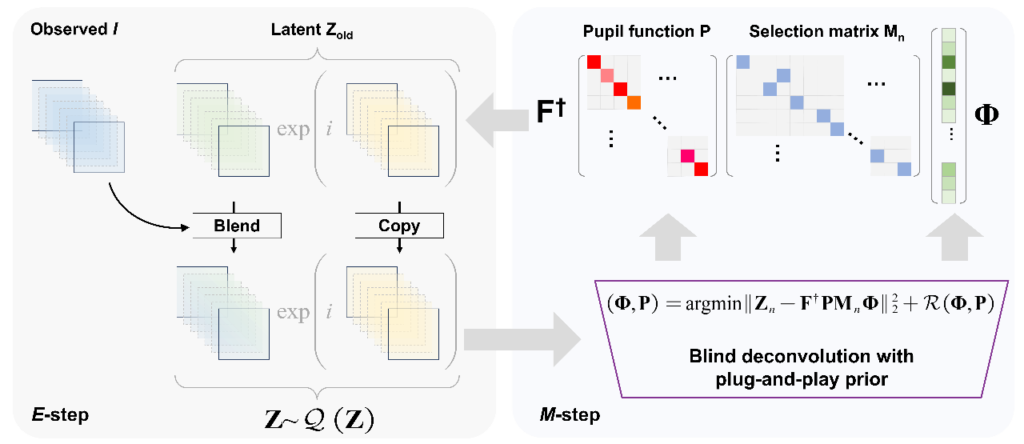
Flowchart for VEM-FPM
What distinguished the VEM-FPM approach is that each raw image was used not only to update the sample wavefront but also to refine the latent variable Z—the complex wavefront that would have arrived at the sensor plane in a perfect, noise-free system. During the E-step, the algorithm estimated Z via proximal gradient descent using intensity inputs and predictions from the current model; in the M-step, the updated Z fields were combined through a quadratic deconvolution to recover both the sample’s Fourier spectrum and the pupil function. The authors found when they applied to H&E-stained rat colon tissue, the method reconstructed an entire 5.3 mm × 5.3 mm field of view without splitting the image into patches or resorting to segment stitching. The reconstruction achieved a synthetic aperture equivalent to a 0.53 NA objective, despite using a physical objective of only NA = 0.08. Compared visually to conventional ePIE and the more robust feature-domain FPM (FD-FPM), the VEM approach eliminated vignetting artifacts and maintained uniform color appearance across the entire field. The intensity profiles extracted from corresponding image rows confirmed that VEM-FPM preserved high-frequency detail where ePIE faltered under the vignetting distortions.
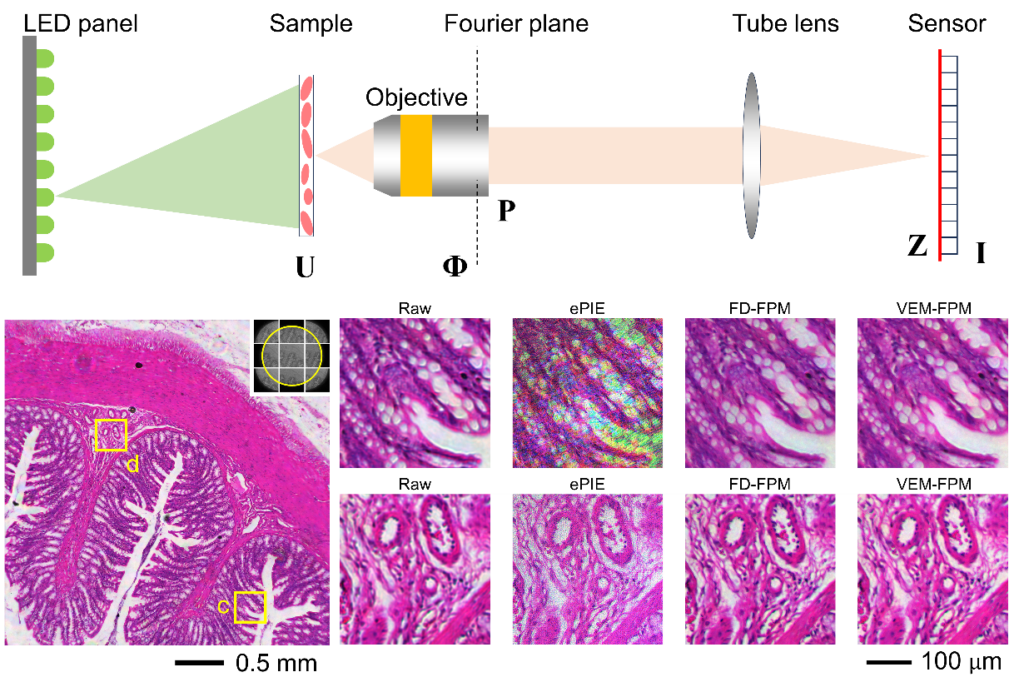
FPM experimental layout and reconstruction results.
The authors also further evaluated the efficiency of their method and found the algorithm processed the full dataset in roughly half the time required by FD-FPM while matching its resolution and outperforming it in producing seamless whole-slide views. A second set of experiments examined quantitative phase retrieval using resolution targets. Here, the method resolved up to 1024 line pairs per millimeter in the high-frequency region, though the lowest phase frequencies—generally irrecoverable in standard FPM—remained only partially reconstructed, consistent with known theoretical limitations. They also deliberately introduced aberrations through sample defocusing. Without prior knowledge of the defocus magnitude or aberration structure, VEM-FPM recovered a sharp wavefront and reconstructed a pupil function whose Zernike components closely matched ground-truth values which suggested that the latent-wavefront strategy naturally compensates for significant phase distortions without external calibration, strengthening the method’s case as a practical tool for large-scale tissue imaging.
In conclusion, Professor Liangcai Cao and colleagues successfully developed a new VEM-FPM method that yields fast, stitching-free reconstructions across very large tissue fields, significantly reducing sensitivity to vignetting and system misalignment. Their new framework outperforms traditional iterative solvers while preserving physical interpretability and robustness. One immediate implication is that high-quality, large-area reconstructions are now attainable with lower-end optics and standard LED arrays. For pathology laboratories—especially those in resource-limited settings—this could lower the barrier to digital whole-slide imaging systems capable of resolving sub-cellular detail. Instead of relying on high-NA objectives and mechanical scanners, laboratories could adopt simplified optical setups that capture large regions at once, leaving the computational model to synthesize high-resolution images. The authors’ demonstration of stitching-free whole-slide reconstruction illustrates how such systems may reduce acquisition time, eliminate alignment errors at tile boundaries, and provide cleaner visualizations for pathologists who increasingly rely on digital workflows augmented by machine learning. Another implication concerns computational tractability. FPM algorithms have historically demanded long processing times because they sought to update the entire sample wavefront via non-convex optimization. Additionally, the VEM framework breaks the problem into two pieces—one non-convex but local and parallelizable, the other convex and closed-form which allow the overall process to accelerate without sacrificing interpretability. From a methodological standpoint, this decomposition bridges physical modeling and statistical inference, offering a structure that may be extended to other imaging modalities where latent wavefields govern the formation of observed intensities.
According to the authors the new technique can be applied in other than stained tissue imaging because phase retrieval sits at the core of many scientific and engineering systems—remote imaging, adaptive optics, holography, and coherent diffractive imaging all grapple with similar inverse problems. Indeed, the VEM framework, with its explicit latent variable and plug-and-play denoising, could serve as a foundation for next-generation solvers that balance model fidelity with algorithmic speed. In that sense, its value extends beyond microscopy, providing a lens through which other inverse problems may be reframed to better handle uncertainties and imperfections in experimental data. For digital pathology specifically, the work aligns with ongoing transitions toward computationally enhanced diagnostics. As machine learning models increasingly analyze whole-slide images for classification, segmentation, and prognostic prediction, the need for consistent and artifact-free slide digitization grows. VEM-FPM produces cleaner, more uniform images, reduces color inconsistencies, and recovers fine structures needed for accurate downstream analysis. The method thus supports a workflow in which optical limitations are resolved computationally and allows diagnostic algorithms to operate on reliable, high-quality visual inputs.
Reference
Shuhe Zhang, Jiayun Li, and Liangcai Cao, “Latent-wavefront Fourier ptychography for stained tissue microscopy,” Photon. Res. 13, 1893-1901 (2025)
Go to Journal of Photon. Res. Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.
Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.

