Significance
Environmental safety and public health are a global concern among various policymakers and stakeholders. Even though great improvement have been made in the treatment and disposal of sludge, a lot has to be done to ensure efficiency as well as human and environment safety. For instance, waste activated sludge provides a favorable environment for the growth of antibiotic resistance genes. Recently, research about the antibiotic resistance genes has attracted significant attention of scientists owing to their potential negative impacts.
Presently, several approaches such as increasing the digestion temperature to thermophilic and prefermentation with alkaline pH have been developed to reduce the antibiotic resistance genes in the sludge. However, most of these methods are not suitable for contaminants such as the metal oxide nanoparticles that are widely used in numerous industries like the cosmetics. This is attributed to the accumulation of the nanoparticles in the sludge which coexisting with the antibiotic resistance genes. The combination results in low performance of the anaerobic digestion. Unfortunately, the effects of the nanoparticles on the distribution of the antibiotic resistance genes during anaerobic digestion of the sludge have not been fully explored.
Waste sludges are generally comprised of complex matrix constituting various microorganisms species. Therefore, the present bacteria are capable of sensing and responding to different environmental changes like the two-component regulatory system in the presence of the stimulus. In a recently published literature, a two-component regulatory system enables bacteria to adapt to different environmental stimuli. To this end, there is a great need to understand the contribution of the two-component regulatory system to antibiotic resistance genes during sludge anaerobic digestion.
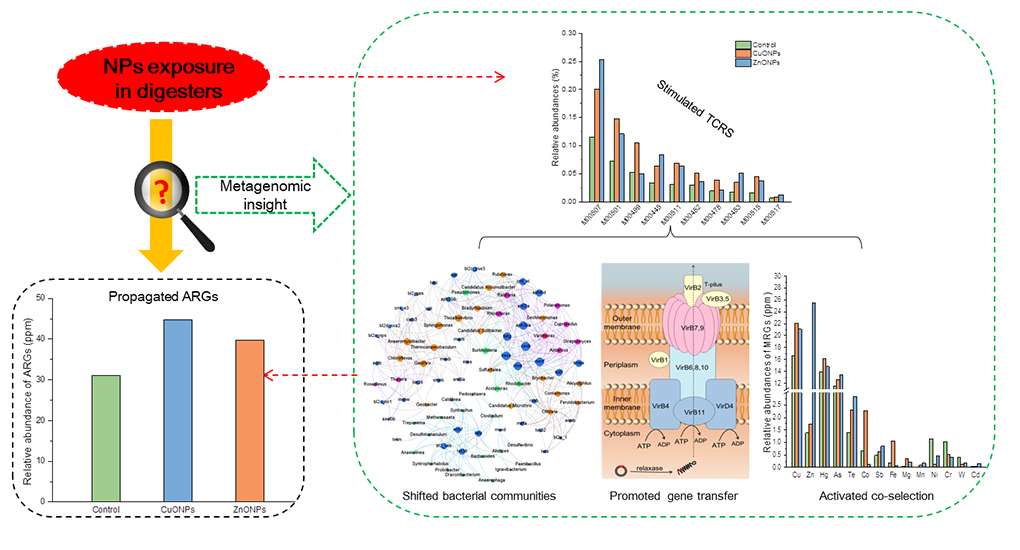
Tongji University researchers: Dr. Xiong Zheng and his colleagues investigated the effects of nanoparticles on the antibiotic resistance genes during sludge anaerobic digestion. They utilized the CuO nanoparticles and ZnO nanoparticles and examined their impacts. Quantitative PCR and metagenomics were used to assess the shifts of the antibiotic resistance genes patterns. They aimed at clarifying the correlation between the triggered two-component regulatory system and altered bacterial taxa. Their work is published in the research journal, Environmental Science: Nano.
From the experimental results, the authors observed that the CuO and ZnO nanoparticles resulted in the propagation of antibiotic resistance genes during sludge anaerobic. However, they had insignificant effects on the resistance mechanisms and types. Consequently, both of the nanoparticles were capable of stimulating the signal transduction and especially the two-component regulatory system that is responsible for pili synthesis, metal tolerance, and quorum sensing.
According to the authors, the study will advance the propagation of antibiotic resistance genes among bacteria in digesters through nanoparticle stimulated signal transduction. Therefore, it provides new insights that enable proper understanding of the antibiotic resistance genes responses to different environmental stimuli while at the same time providing a reference for future studies. This will lead to effective treatment of sludge wastes with a key interest in environmental and human safety.
Reference
Huang, H., Chen, Y., Yang, S., & Zheng, X. (2019). CuO and ZnO nanoparticles drive the propagation of antibiotic resistance genes during sludge anaerobic digestion: possible role of stimulated signal transduction. Environmental Science: Nano, 6(2), 528-539.
Go To Environmental Science: Nano Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.
Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.