Significance
Self-assembly and self-organisation are exploited across virtually all areas of nanoscience and nanoengineering as a means of patterning matter, often for subsequent application in devices, sensors, or actuators. Arguably, the archetypal example of a self-organising nanostructured system is a sample of colloidal nanoparticles suspended (or dissolved) in a surrounding solvent: nanoparticle-solvent films deposited on solid substrates are associated with a rich dynamic behaviour which gives rise to a wide variety of striking self-organized patterns.
In the far-from-equilibrium regime (i.e. where solvent evaporation is rapid), a remarkably broad array of intricate, spatially correlated patterns form including “foam-like” cellular networks, labyrinthine structures similar to those formed in spinodal decomposition of binary fluids, and fractal morphologies [1-4]. In many ways the system is a playground for self-organisation driven far from equilibrium (and, coincidentally, has many parallels with the physics of coffee stains). Its ability to generate a panoply of patterns across a wide range of length-scales provides a stringent test of the ability of machine learning algorithms to sift and classify self-organised and self-assembled structures.
In order to map (and modify) the structured of self-organised systems, scanning probe microscopes (SPMs) are very often the tool of choice. SPMs are conceptually straight-forward (but rather more difficult to realise experimentally): a sharp probe – often terminated by a single atom – is scanned back and forth across a surface while interactions between the tip and the sample are measured. These tip-sample forces have a number of potential origins: electrostatic, chemical, magnetic, and van der Waals interactions among them..
Currently, researchers spend significant time manually searching through large volumes of data produced during scanning probe imaging to identify specific patterns and motifs formed via self-assembly and self-organization. As such, there is need that this cumbersome approach be simplified. In this regard, researchers from the University of Nottingham — Oliver Gordon (PhD candidate), Dr. Jo Hodgkinson and Professor Philip Moriarty — together with Dr. Steff Farley and Dr. Eugenie Hunsicker at Loughborough University in the UK developed an innovative combination of Monte Carlo simulations, general statistics, and machine learning to automatically distinguish several spatially correlated patterns in a mixed, highly varied data set of real AFM images of self-organized nanoparticles. Their work is currently published in the research journal, Nano Letters.
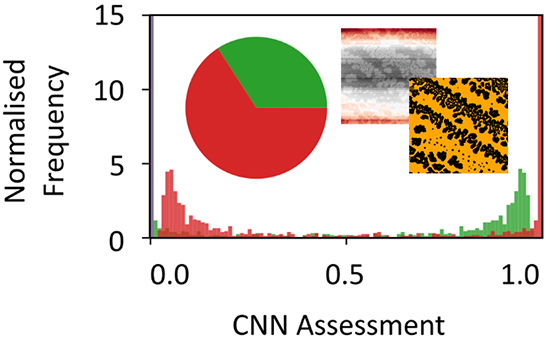
In their approach, they focused on demonstrating that it is possible to use simulated AFM images of self-organized nanoparticle assemblies as automatically labeled training data for a neural network, which then correctly generalizes to real AFM scans. As such, they employed an optimized preprocessing routine for real images of these specific structures, which was then combined with a denoising autoencoder to provide effective, automated binarization of real images. The researchers then combined these systems to quickly find experimental AFM images of these specific structures in a data set of 5519 scans of multiple structures and surfaces collected at a wide range of scan qualities and sizes from 500 nm to 90 μm.
The authors showed that it is possible to use simple simulations to correctly find specific structures in a data set of mixed AFM images without the need for manually labeled training data. In addition, the reported method was proven to be heavily reliant on good preprocessing and binarization. While noise removal was also particularly vital, a denoising autoencoder trained on the simulated data performed extremely well at this task and may be applicable elsewhere to smooth features and remove processing artifacts.
In summary, the study demonstrated the development of an automated identification method for nanostructured patterns. The method was proven as an effective first stage of file search, capable of isolating files and locations of interest. and significantly reduces the time required to search data sets.s. Provided that the structures of interest can be simulated, the strategy and protocols they describe could be easily adapted to other self-organized systems and data sets.
Reference
Oliver M. Gordon, Jo E. A. Hodgkinson, Steff M. Farley, Eugenie L. Hunsicker, and Philip J. Moriarty. Automated Searching and Identification of Self-Organized Nanostructures. Nano Letters 2020; volume 20, page 7688−7693.
 Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.
Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.