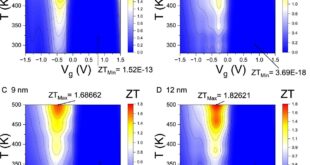
Significance
Ideally, a nanopore is a nanoscale opening in a membrane that separates two fluidic chambers. At present, nanopores have become powerful tools for performing biological analysis, enabling label-free, low-cost sensing of single biomolecules using a purely electrical sensing approach. Technically, a voltage-clamp circuit supplies a voltage bias to capture and pass individual charged molecules, such as DNA, from one chamber to the other. So far, controlling translocation has remained the main snag in the nanopore field. It is well documented that solid-state nanopores lack a translocation control mechanism that can slow-down the DNA while ensuring a high sensing voltage to generate ample signal-to-noise ratio (SNR) for robust feature detection. To this resolve, recent publications have reported on a new class of “two-pore” devices that enables simultaneous capture of a single DNA molecule by two closely spaced pores. Two-pore devices, in principle, enable independent control of the DNA sensing bias and the bias driving DNA translocation. Regardless, current solid-state nanopore technology still lacks an effective means for combined conformational and dynamic control of translocating single molecules.
Therefore, methods for reducing and directly controlling the speed of DNA through a nanopore are urgently needed to enhance sensing performance for direct strand sequencing and detection/mapping of sequence-specific features. In this context, a group of researchers from the Ontera Incorporation in California: Dr. Xu Liu, Dr. Roland Nagel and Dr. William B. Dunbar together with Dr. Yuning Zhang and Professor Walter Reisner from the Department of Physics at McGill University in Montreal developed a novel method that uses two independently controllable nanopores operated with an active control logic for reducing and controlling the speed of DNA. To be precise, the researchers proposed to construct a dual pore device that employed a Field-Programmable Gate Array (FPGA) to implement active feed-back control. Their work is currently published in the research journal, Small.
Briefly, in their approach, the pores were positioned sufficiently close so as to permit co-capture of a single DNA by both pores. Once co-capture occurred, control logic turned on constant competing voltages at the pores leading to a “tug-of-war” whereby opposing forces were applied to regions of the molecules threading through the pores. The forces exerted both conformational and speed control over the co-captured molecule, removing folds and reducing the translocation rate.
The authors found that the mean tug-of-war duration as a function of reversed bias exhibited a resonance like structure, with the peak lifetime corresponding to a balance of the opposing forces and a physical scenario whereby DNA motion was controlled entirely by diffusional sliding between the pores. Additionally, they reported that the diffusion process was in fact sub-diffusive, ensuing a greater number of longtime tug-of-war states than would be predicted on the basis of simple diffusion.
In summary, the research team demonstrated the application of a two-pore advanced technology for reducing and controlling the speed of DNA that uses two independently controllable nanopores operated with an active control logic. The study was inspired by the fact that a two-pore tug-of-war implemented with active feedback gives rise to predominantly linearized translocation signatures, obviating the folding; a shortfall that offers significant challenge in electrical barcoding approaches. Overall, the researchers demonstrated that through their new approach, one could detect correlated motion of bound streptavadin tags at pores 1 and 2 on molecules undergoing two-pore tug-of-war, thereby indicating enhanced detection of the tags over a single-pore measurement of the same molecules and the ability to use physical tags to independently assess translocation velocity.
“Active and feedback control has a long history of enabling innovations, and we’re excited to be leading the combined use of nanotechnology and control methods to enable rapid and thorough interrogation of single molecules, for a variety of use cases. This can enable long-sequencing, as well as mapping genomic and genome-associated features and feature modifications, which is quite relevant for epigenetics analysis.” Said Dr. William B. Dunbar in a statement to Advance in Engineering.
Professor Walter Reisner commented on the collaborative research work: “There have been quite spectacular advances made in solid-state pore technology over the past five years, but achieving consistently linearized and slowed-down translocation is still a major challenge in the solid-state nanopore field. I find two pore tug-of-war control with active logic to be an extremely elegant approach to this problem. It is conceptually simple and involves a minimum increase in the complexity of a standard single-pore experiment, which should make it more scalable relative to competing approaches. In particular, the ability to implement active control over a single-molecule translocation could be an extremely powerful capability, leading to radical advances such as the ability to interrogate targeted regions of large molecules and counteract Brownian fluctuations so that thermal noise can be reduced.”
Reference
Xu Liu, Yuning Zhang, Roland Nagel, Walter Reisner, William B. Dunbar. Controlling DNA Tug-of-War in a Dual Nanopore Device. Small 2019, volume 15, page 1901704.
Go To Small Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.
Advances in Engineering Advances in Engineering features breaking research judged by Advances in Engineering advisory team to be of key importance in the Engineering field. Papers are selected from over 10,000 published each week from most peer reviewed journals.